Thought Chart: Tracking Dynamic EEG Brain Connectivity with Unsupervised Manifold Learning
October 13th, 2016
Authors
Xing, M., Ajilore, O., Wolfson, O., Abbott, C., MacNamaram, A., Tadayonnejad, R., Forbes, A., Phan, K.L., Klumpp, H., Leow, A.About
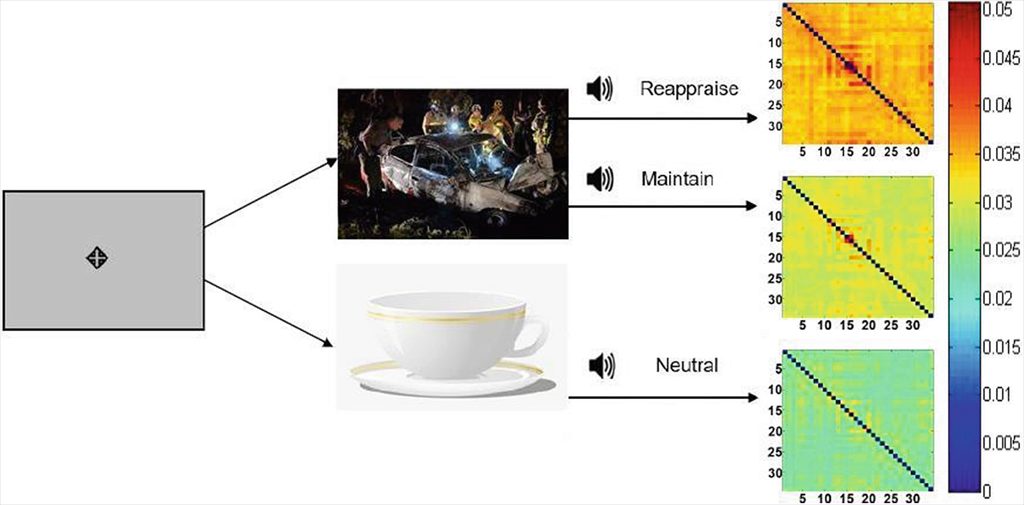
Assuming that the topological space containing all possible brain states forms a very high-dimensional manifold, this paper proposes an unsupervised manifold learning framework to reconstruct and visualize this manifold using EEG brain connectivity data acquired from a group of healthy volunteers. Once this manifold is constructed, the temporal sequence of an individual’s EEG activities can then be represented as a trajectory or thought chart in this space. Our framework first applied graph dissimilarity space embedding to the temporal EEG connectomes of 20 healthy volunteers, both at rest and during an emotion regulation task (ERT), followed by local neighborhood reconstruction then nonlinear dimensionality reduction (NDR) in order to reconstruct and embed the learned manifold in a lower-dimensional Euclidean space. We showed that resting and ERT thought charts represent distinct trajectories, and that the manifold resembles dynamical systems on the torus. Additionally, new trajectories can be inserted on-line via out-of-sample embedding, thus providing a novel data-driven framework for classifying brain states, with potential applications in neurofeedback via real-time thought chart visualization.
Keywords: Thought chart; Graph dissimilarity embedding; Nonlinear dimensionality reduction; EEG connectome; Emotion regulation
DOI: 10.1007/978-3-319-47103-7_15
Resources
Citation
Xing, M., Ajilore, O., Wolfson, O., Abbott, C., MacNamaram, A., Tadayonnejad, R., Forbes, A., Phan, K.L., Klumpp, H., Leow, A., Thought Chart: Tracking Dynamic EEG Brain Connectivity with Unsupervised Manifold Learning, Brain Informatics and Health: International Conference, BIH 2016, Omaha, NE, Springer international Publishing, pp. 149-157, October 13th, 2016.