FixingTIM: Interactive Exploration of Sequence and Structural Data to Identify Functional Mutations in Protein Families
August 1st, 2014
Categories: Applications, Visualization
Authors
Luciani, T., Wenskovitch, J., Chen, K., Koes, D., Travers, T., Marai, G.E.About
Background: Knowledge of the 3D structure and functionality of proteins can lead to insight into the associated cellular processes, speed up the creation of pharmaceutical products, and develop drugs that are more effective in combating disease.
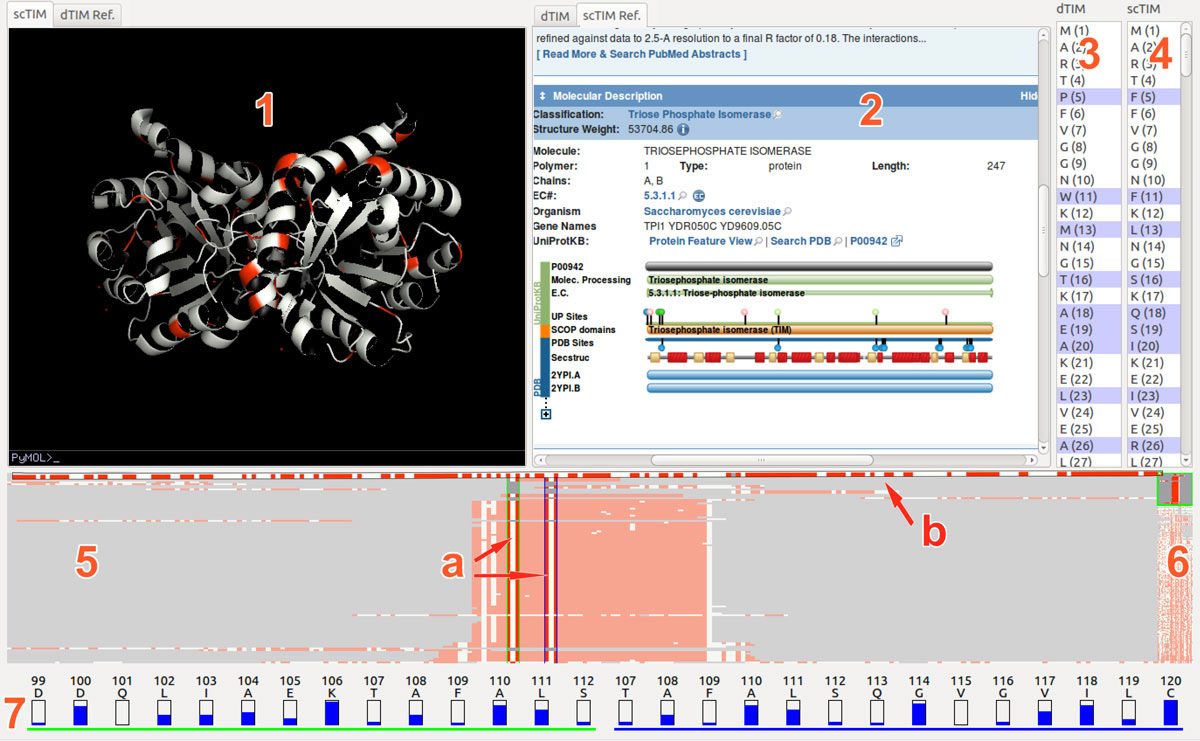
Methods: We present the design and implementation of a visual mining and analysis tool to help identify protein mutations across a family of structural models and to help discover the effect of these mutations on protein function. We integrate 3D structure and sequence information in a common visual interface; multiple linked views and a computational backbone allow comparison at the molecular and atomic levels, while a novel trend-image visual abstraction allows for the sorting and mining of large collections of sequences and of their residues.
Results: We evaluate our approach on the triosephosphate isomerase (TIM) family structural models and sequence data and show that our tool provides an effective, scalable way to navigate a family of proteins, as well as a means to inspect the structure and sequence of individual proteins.
Conclusions: The TIM application shows that our tool can assist in the navigation of families of proteins, as well as in the exploration of individual protein structures. In conjunction with domain expert knowledge, this interactive tool can help provide biophysical insight into why specific mutations affect function and potentially suggest additional modifications to the protein that could be used to rescue functionality.
Resources
Citation
Luciani, T., Wenskovitch, J., Chen, K., Koes, D., Travers, T., Marai, G.E., FixingTIM: Interactive Exploration of Sequence and Structural Data to Identify Functional Mutations in Protein Families, BMC Proceedings 2014, vol 8, 8(Suppl 2):S3, pp. 1-0, August 1st, 2014.