Science Grid This Week Features EVL Alumni Research
March 29th, 2006
Categories: Applications, Software, Supercomputing, Virtual Medicine, Visualization
About
EVL alumni and Argonne National Laboratory staff members, Mike Papka’s and Joe Insley’s research efforts and collaborative work in the development of arterial simulations / visualization are featured in “Science Grid This Week”.
Grid Gets the Blood Flowing
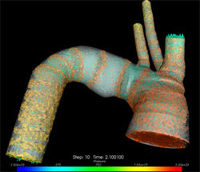
A collaboration of mathematicians, middleware developers and visualization scientists has demonstrated the most comprehensive three-dimensional model of human arterial blood flow ever attempted. The simulation of the human arterial tree, the network of arteries throughout the human body, was completed using TeraGrid resources.
“We want to be able to study diseases like atherosclerosis in more detail,” said George Karniadakis from Brown University. “What’s the effect on the arteries in the brain if there’s atherosclerosis, or plaque buildup, in the carotid artery? Individual arteries have been studied by many research groups, but they have to specify artificial conditions at the edges of each artery.” With a global blood circulation model such guesses at conditions are not necessary, and scientists can see the effect of a disturbance in blood flow anywhere in the body. Such simulations may impact many areas of medical research.
Over the past 10 years, Karniadakis and his research group developed a computer program that simulates the largest 55 arteries and 27 bifurcations, or places where an artery splits into two, in the tree. Until last year, the simulations were run on only one computing resource at a time, modeling at most two or three branching sites at once. But in early 2005, the collaboration proposed harnessing the combined power of the TeraGrid’s many computing resources to simulate, and visualize in real time, more than a dozen branching sites?
More www.interactions.org/sgtw/2006/0329/arterial_tree_more.html